Sklearn Random Forest Classification
SKLearn Classification using a Random Forest Model
import platform
import sys
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import matplotlib
matplotlib.style.use('ggplot')
%matplotlib inline
import time
from scipy.stats import randint as sp_randint
import seaborn as sns
import sklearn
from sklearn.preprocessing import OneHotEncoder
from sklearn.ensemble import RandomForestClassifier
from sklearn.mixture import GaussianMixture
from sklearn.model_selection import RandomizedSearchCV, GridSearchCV
from sklearn.model_selection import cross_val_score, train_test_split
from sklearn import metrics
from sklearn.metrics import roc_auc_score, roc_curve, auc, classification_report
from sklearn.metrics import f1_score, precision_score, recall_score
from sklearn.metrics import mean_squared_error, cohen_kappa_score, make_scorer
from sklearn.metrics import confusion_matrix, accuracy_score, average_precision_score
from sklearn.metrics import precision_recall_curve, SCORERS
from sklearn.model_selection import learning_curve
from sklearn.model_selection import ShuffleSplit
from sklearn.externals import joblib
from operator import itemgetter
from tabulate import tabulate
Here is the environment for this notebook
print('Operating system version....', platform.platform())
print("Python version is........... %s.%s.%s" % sys.version_info[:3])
print('scikit-learn version is.....', sklearn.__version__)
print('pandas version is...........', pd.__version__)
print('numpy version is............', np.__version__)
print('matplotlib version is.......', matplotlib.__version__)
Operating system version.... Windows-10-10.0.16299-SP0
Python version is........... 3.6.4
scikit-learn version is..... 0.19.1
pandas version is........... 0.22.0
numpy version is............ 1.13.3
matplotlib version is....... 2.1.2
Define a TIme Class to computer total execution time
class Timer:
def __init__(self):
self.start = time.time()
def restart(self):
self.start = time.time()
def get_time(self):
end = time.time()
m, s = divmod(end - self.start, 60)
h, m = divmod(m, 60)
time_str = "%02d:%02d:%02d" % (h, m, s)
return time_str
Define a correlation plot method
def Correlation_plot(df):
plt.ioff()
red_green = ["#ff0000", "#00ff00"]
sns.set_palette(red_green)
np.seterr(divide='ignore', invalid='ignore')
g = sns.pairplot(df,
diag_kind = 'kde',
hue = 'Altitude',
markers = ["o", "D"],
size = 1.5,
aspect = 1,
plot_kws = {"s": 6})
g.fig.subplots_adjust(right = 0.9)
plt.show()
Define a method to load the Bottle Rocket Data Set
We load the Bottle Rocket data into two datasets: train and test. Since we are doing cross-validation, we only need the train dataset to do training. The test dataset is our “out-of-sample” data that will be used only after training is done. This is an attempt to simulate a production environment.
def LoadData():
global X_train, y_train, X_test, y_test, train, test
global feature_columns, response_column, n_features
model_full = pd.read_csv("C:/sm/BottleRockets/model-8-1.csv")
response_column = ['Altitude']
feature_columns = ['BoxRatio','Thrust', 'Velocity', 'OnBalRun', 'vwapGain']
n_features = len(feature_columns)
mask = feature_columns + response_column
model = model_full[mask]
print('Model dataset:\n', model.head(5))
print('\nDescription of model dataset:\n', model[feature_columns].describe(include='all'))
Correlation_plot(model)
# Split the data into Train and Test with Train having 80% and test 20% each
train_full, test_full = np.split(model_full.sample(frac=1), [int(.8*len(model))])
X_train = train_full[feature_columns].as_matrix()
y_train_ = train_full[response_column].as_matrix()
X_test = test_full[feature_columns].as_matrix()
y_test_ = test_full[response_column].as_matrix()
y_train = np.reshape(y_train_, len(y_train_))
y_test = np.reshape(y_test_, len(y_test_))
train = train_full[mask]
test = test_full[mask]
print('Shape of train: ', train.shape)
print('Shape of test: ', test.shape)
return
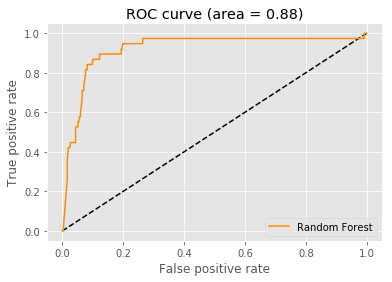
Define a method to plot a ROC Curve
def ROC_Curve(rf, auc):
one_hot_encoder = OneHotEncoder()
rf_fit = rf.fit(X_train, y_train)
fit = one_hot_encoder.fit(rf.apply(X_train))
y_predicted = rf.predict_proba(X_test)[:, 1]
false_positive, true_positive, _ = roc_curve(y_test, y_predicted)
plt.figure()
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(false_positive, true_positive, color='darkorange', label='Random Forest')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.title('ROC curve (area = %0.2f)' % auc)
plt.legend(loc='best')
plt.show()
Define a method to print the model performance
def Print_Metrics(saved_rf):
print('\nModel performance on the test data set:')
# print('Train Accuracy.......', accuracy_score(y_train, best_model.predict(X_train)))
# print('Validate Accuracy....', accuracy_score(y_valid, best_model.predict(X_valid)))
y_predict_test = best_model.predict(X_test)
mse = metrics.mean_squared_error(y_test, y_predict_test)
logloss_test = metrics.log_loss(y_test, y_predict_test)
accuracy_test = metrics.accuracy_score(y_test, y_predict_test)
accuracy_test2 = best_model.score(X_test, y_test)
F1_test = metrics.f1_score(y_test, y_predict_test)
precision_test = precision_score(y_test, y_predict_test, average='binary')
precision_test2 = metrics.precision_score(y_test, y_predict_test)
recall_test = recall_score(y_test, y_predict_test, average='binary')
auc_test = metrics.roc_auc_score(y_test, y_predict_test)
r2_test = metrics.r2_score(y_test, y_predict_test)
#test_auc = h2o.get_model("best_rf").model_performance(test_data=test).auc()
#print('Best model performance based on auc: ', test_auc)
header = ["Metric", "Test"]
table = [
["logloss", logloss_test],
["accuracy", accuracy_test],
["precision", precision_test],
["F1", F1_test],
["r2", r2_test],
["AUC", auc_test]
]
print(tabulate(table, header, tablefmt="fancy_grid"))
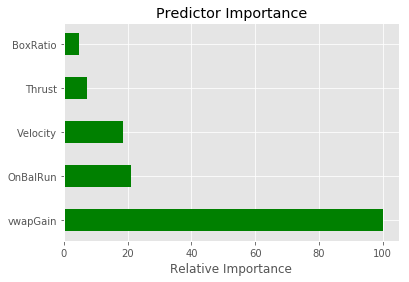
Define a method to plot the predictor importance
def Plot_predictor_importance(best_model, feature_columns):
feature_importances = best_model.feature_importances_
sorted_idx = np.argsort(feature_importances)
y_pos = np.arange(sorted_idx.shape[0]) + .5
fig, ax = plt.subplots()
ax.barh(y_pos,
feature_importances[sorted_idx],
align='center',
color='green',
ecolor='black',
height=0.5)
ax.set_yticks(y_pos)
ax.set_yticklabels(feature_columns)
ax.invert_yaxis()
ax.set_xlabel('Relative Importance')
ax.set_title('Predictor Importance')
plt.show()
Define a utility function to report best scores
def Report_scores(results, n_top=3):
for i in range(1, n_top + 1):
candidates = np.flatnonzero(results['rank_test_score'] == i)
for candidate in candidates:
print("Model with rank: {0}".format(i))
print("Mean validation score: {0:.3f} (std: {1:.3f})".format(
results['mean_test_score'][candidate],
results['std_test_score'][candidate]))
print("Parameters: {0}".format(results['params'][candidate]))
print("")
Define a method to print the Confusion Matrix and the performance metrics
def Print_confusion_matrix(cm, auc, heading):
print('\n', heading)
print(cm)
true_negative = cm[0,0]
true_positive = cm[1,1]
false_negative = cm[1,0]
false_positive = cm[0,1]
total = true_negative + true_positive + false_negative + false_positive
accuracy = (true_positive + true_negative)/total
precision = (true_positive)/(true_positive + false_positive)
recall = (true_positive)/(true_positive + false_negative)
misclassification_rate = (false_positive + false_negative)/total
F1 = (2*true_positive)/(2*true_positive + false_positive + false_negative)
print('accuracy.................%7.4f' % accuracy)
print('precision................%7.4f' % precision)
print('recall...................%7.4f' % recall)
print('F1.......................%7.4f' % F1)
print('auc......................%7.4f' % auc)
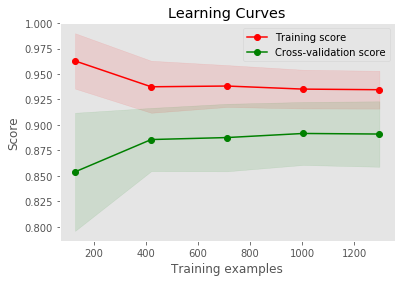
Plot the learning curves
def Plot_learning_curve(estimator, title, X, y, ylim = None, cv = None,
n_jobs = 1, train_sizes = np.linspace(0.1, 1.0, 5)):
plt.figure()
plt.title(title)
if ylim is not None:
plt.ylim(*ylim)
plt.xlabel("Training examples")
plt.ylabel("Score")
train_sizes, train_scores, test_scores = learning_curve(estimator,
X, y,
cv = cv,
n_jobs = n_jobs,
train_sizes = train_sizes)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
plt.grid()
plt.fill_between(train_sizes, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.1,
color="r")
plt.fill_between(train_sizes, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1, color="g")
plt.plot(train_sizes, train_scores_mean, 'o-', color="r",
label="Training score")
plt.plot(train_sizes, test_scores_mean, 'o-', color="g",
label="Cross-validation score")
plt.legend(loc="best")
plt.show()
return
Define the hyperparameters for a random search
def Random_Search():
global best_model, saved_moldel
param_grid = {"n_estimators": range(20, 100, 2),
"max_depth": range(4, 50, 2),
"min_samples_leaf": range(2, 100, 2),
"max_features": sp_randint(1, n_features),
"min_samples_split": sp_randint(2, 10),
"bootstrap": [True, False],
"criterion": ["gini", "entropy"]}
clf = RandomForestClassifier(class_weight = 'balanced')
n_iter_search = 500
estimator = RandomizedSearchCV(clf,
param_distributions = param_grid,
n_iter = n_iter_search,
scoring = 'roc_auc',
verbose = 0,
n_jobs = 1)
fit = estimator.fit(X_train, y_train)
# Cross validation with 20 iterations to get smoother mean test and train
# score curves, each time with 20% data randomly selected as a validation set.
cv_ = ShuffleSplit(n_splits = 20, test_size = 0.20, random_state = 0)
Plot_learning_curve(estimator,
'Learning Curves',
X_train, y_train,
cv = cv_,
n_jobs = 1)
Report_scores(estimator.cv_results_, n_top = 3)
best_model = estimator.best_estimator_
print('\nbest_model:\n', best_model)
print('\nFeature Importances:', best_model.feature_importances_)
Plot_predictor_importance(best_model, feature_columns)
y_predicted = best_model.predict(X_train)
probabilities = best_model.predict_proba(X_train)
c_report = classification_report(y_train, y_predicted)
print('\nClassification report:\n', c_report)
y_predicted_train = best_model.predict(X_train)
cm = confusion_matrix(y_train, y_predicted_train)
auc = roc_auc_score(y_train, y_predicted_train)
Print_confusion_matrix(cm, auc, 'Confusion matrics of the training dataset')
y_predicted = best_model.predict(X_test)
cm = confusion_matrix(y_test, y_predicted)
auc = roc_auc_score(y_test, y_predicted)
ntotal = len(y_test)
correct = y_test == y_predicted
numCorrect = sum(correct)
percent = round( (100.0*numCorrect)/ntotal, 6)
print("\nCorrect classifications on test data: {0:d}/{1:d} {2:8.3f}%".format(numCorrect, ntotal, percent))
prediction_score = 100.0*best_model.score(X_test, y_test)
print('Random Forest Prediction Score on test data: %8.3f' % prediction_score)
model_path = 'C:/sm/BottleRockets/trained_models/sklearn_rf_classify.pkl'
joblib.dump(best_model, model_path)
saved_model = joblib.load(model_path)
y_predicted_test = best_model.predict(X_test)
cm = confusion_matrix(y_test, y_predicted_test)
auc = roc_auc_score(y_test, y_predicted_test)
Print_confusion_matrix(cm, auc, 'Confusion matrics of the test dataset')
ROC_Curve(best_model, auc)
Print_Metrics(saved_model)
return
Run the random search
my_timer = Timer()
LoadData()
Random_Search()
elapsed = my_timer.get_time()
print("\nTotal compute time was: %s" % elapsed)
Model dataset:
BoxRatio Thrust Velocity OnBalRun vwapGain Altitude
0 0.166 0.166 0.317 0.455 -0.068 0
1 0.071 0.068 0.170 0.482 -0.231 0
2 -0.031 -0.031 0.109 0.531 0.115 0
3 -0.186 -0.193 0.344 0.548 0.111 0
4 -0.147 -0.147 0.326 0.597 0.157 1
Description of model dataset:
BoxRatio Thrust Velocity OnBalRun vwapGain
count 2025.000000 2025.000000 2025.000000 2025.000000 2025.000000
mean 1.193801 1.048527 0.282673 2.422911 0.522294
std 2.909786 2.015741 0.919509 2.247012 1.218212
min -0.938000 -0.938000 -0.816000 0.455000 -0.734000
25% 0.183000 0.184000 -0.021000 1.317000 0.098000
50% 0.470000 0.511000 -0.009000 1.845000 0.240000
75% 1.143000 1.227000 -0.002000 2.713000 0.523000
max 56.686000 37.219000 11.064000 41.696000 31.841000

Shape of train: (1620, 6)
Shape of test: (405, 6)
Model with rank: 1
Mean validation score: 0.897 (std: 0.023)
Parameters: {'bootstrap': True, 'criterion': 'entropy', 'max_depth': 28, 'max_features': 3, 'min_samples_leaf': 68, 'min_samples_split': 4, 'n_estimators': 22}
Model with rank: 2
Mean validation score: 0.895 (std: 0.018)
Parameters: {'bootstrap': True, 'criterion': 'gini', 'max_depth': 38, 'max_features': 2, 'min_samples_leaf': 46, 'min_samples_split': 2, 'n_estimators': 20}
Model with rank: 3
Mean validation score: 0.895 (std: 0.021)
Parameters: {'bootstrap': True, 'criterion': 'gini', 'max_depth': 32, 'max_features': 1, 'min_samples_leaf': 28, 'min_samples_split': 2, 'n_estimators': 68}
best_model:
RandomForestClassifier(bootstrap=True, class_weight='balanced',
criterion='entropy', max_depth=28, max_features=3,
max_leaf_nodes=None, min_impurity_decrease=0.0,
min_impurity_split=None, min_samples_leaf=68,
min_samples_split=4, min_weight_fraction_leaf=0.0,
n_estimators=22, n_jobs=1, oob_score=False, random_state=None,
verbose=0, warm_start=False)
Feature Importances: [ 0.03158615 0.0482742 0.65863164 0.12245056 0.13905746]
Classification report:
precision recall f1-score support
0 0.97 0.91 0.94 1433
1 0.53 0.79 0.63 187
avg / total 0.92 0.89 0.90 1620
Confusion matrics of the training dataset
[[1301 132]
[ 40 147]]
accuracy................. 0.8938
precision................ 0.5269
recall................... 0.7861
F1....................... 0.6309
auc...................... 0.8470
Correct classifications on test data: 367/405 90.617%
Random Forest Prediction Score on test data: 90.617
Confusion matrics of the test dataset
[[335 32]
[ 6 32]]
accuracy................. 0.9062
precision................ 0.5000
recall................... 0.8421
F1....................... 0.6275
auc...................... 0.8775
Model performance on the test data set:
╒═══════════╤═══════════╕
│ Metric │ Test │
╞═══════════╪═══════════╡
│ logloss │ 3.4113 │
├───────────┼───────────┤
│ accuracy │ 0.901235 │
├───────────┼───────────┤
│ precision │ 0.484848 │
├───────────┼───────────┤
│ F1 │ 0.615385 │
├───────────┼───────────┤
│ r2 │ -0.161623 │
├───────────┼───────────┤
│ AUC │ 0.874731 │
╘═══════════╧═══════════╛
Total compute time was: 04:19:47
Discussion
That’s not bad, for our first try. Our Bottle Rocket dataset is unbalanced (it’s 8:1). Sci-kit Learn has an argument to it’s models: class_weight = ‘balanced’. This helps with a unbalanced dataset. This is a great advantage over TensorFlow’s high-level API (random_forest.TensorForestEstimator). There is no such argument to help with unbalanced datasets.
We could not wait to use these results. HedgeTools now has a option to use the trained model (sklearn_rf_classify.pkl). It’s not good enough for “program trading”, but when we combine the model with an experienced day-trader, the results are very encouraging.